Single-cell multi-omics analysis of pr CD4+ cells from individuals in the POISED study
The POISED trial design3 is illustrated in Fig. 1a. Participants were randomly assigned to placebo or active treatment groups to be built up and maintained on 4,000 mg peanut protein until week 104 then discontinued peanut dosing (peanut-0 group) or given 300 mg peanut protein daily (peanut-300 group) after week 104. DBPCFCs to 4,000 mg peanut protein were done at BL and week 104 for all participants and at weeks 117, 130, 143 and 156 for participants who passed the DBPCFC at the last time point. Excluding 16 participants who dropped out of the study, 80 of 81 participants in the active treatment group passed the DBPCFC at week 104 and were desensitized, whereas only one of 23 participants in the placebo group passed the week 104 DBPCFC. Twenty-one of 51 (41.2%) peanut-0 group desensitized participants achieved SU, defined as passing DBPCFC at week 117 after 13 weeks of peanut avoidance after OIT. The remaining 30 participants in the peanut-0 group who passed DBPCFC at week 104 but failed at week 117, were defined as only desensitized.
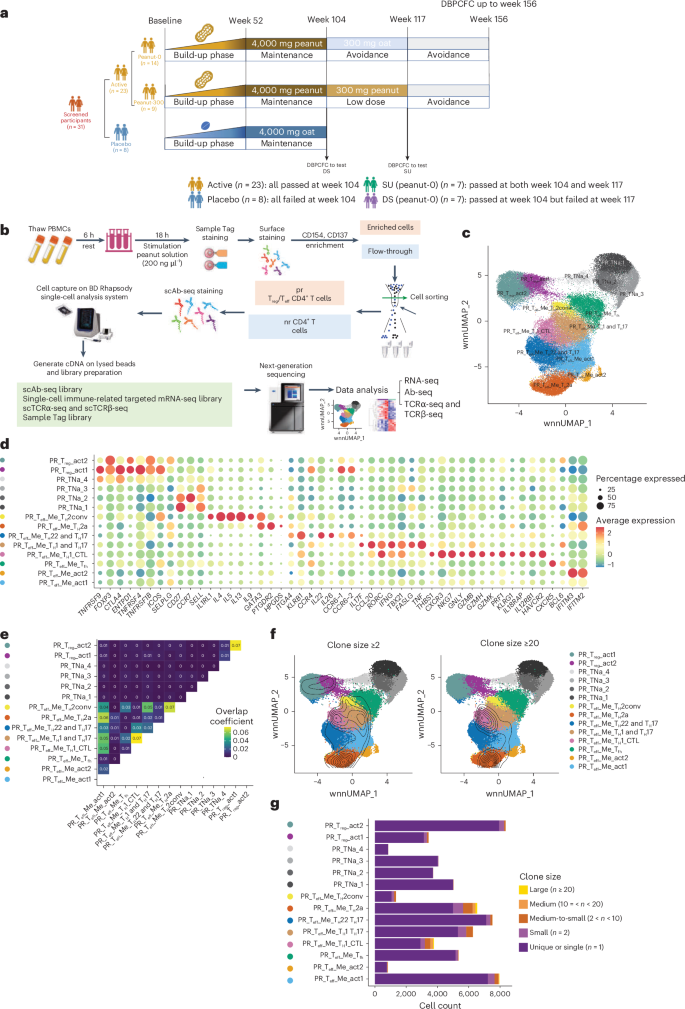
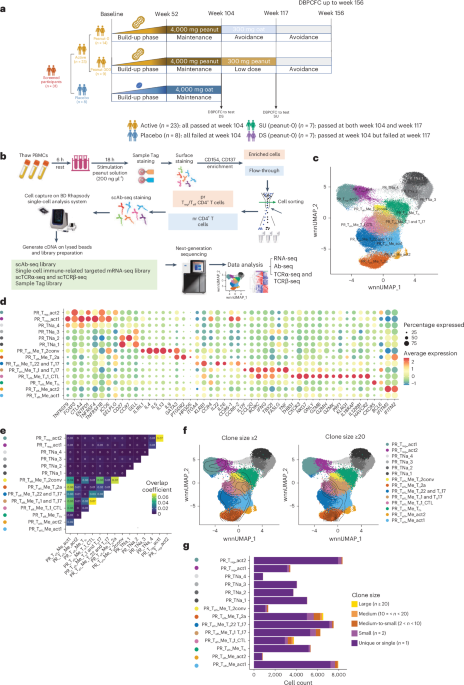
a, Design of the POISED study and sample number in this single-cell multi-omics analysis. Analyses in this study focused on tests of DS (based on the active and placebo groups at BL and week 104) and SU (based on the peanut-0 group at BL, week 104 and week 117). The peanut-300 post-OIT samples and samples from other time points were primarily used in the analyses of tissue-resident T cell clones. b, Schematic of the sample processing workflow. c, UMAP of 72,491 pr CD4+ cells obtained from 31 participants of the POISED trail at several time points and 5 controls without allergy, colored according to cell phenotype base on both scRNA-seq and scAb-seq data integrated using WNN. d, Dot plots of selected marker genes for each cell phenotype displaying the average expression and frequency of expression for each gene. e, Heatmap of clonal overlap rate between each subcluster pairs. f, Clone overlay of the position of clonally expanded cells with clone size large or equal to 2 or 20 mapped onto the corresponding UMAPs. The topographic (contour) lines represent areas of similar density of clonally expanded cells on UMAP, with tighter lines indicating regions of higher density. g, Stacked bar plot showing the cell counts of each clonal size group in each cell subcluster.
Source data
In this trial, we analyzed peripheral blood mononuclear cells (PBMCs) from 31 participants of POISED, selected for having complete time course sampling and sufficient cell numbers available: 23 active participants, who underwent OIT and passed DBPCFC at week 104 (14 peanut-0 participants, of whom seven passed (SU) and seven failed DS at week 117, and nine peanut-300 participants, of whom five passed and four failed DBPCFC at week 117); eight placebo participants; and five age-matched and sex-matched controls without PA (Fig. 1a and Supplementary Tables 1 and 2). We stimulated PBMCs with peanut solution for 18 h and sorted pr CD4+ T cells (CD4+CD137+ or CD4+CD154+) and non-reactive (nr) CD4+ T cells (CD4+CD154−CD137−)9,16. Sorted cells were processed using the BD Rhapsody platform for single-cell RNA sequencing (scRNA-seq) with a focus on 432 immune-relevant target transcripts (399 commercially predefined target panel complemented with 33 targets of interest; Supplementary Table 3) and 42 immune-relevant single-cell antibody sequencing (scAb-seq) markers for protein level detection (Supplementary Table 4), as well as single-cell paired TCRα and TCRβ sequencing (Fig. 1b).
After quality control and removal of multiplets and undetermined cells, we recovered 160,336 cells with high-quality sequencing data. The resulting transcriptomics and protein expression data were separately processed using reciprocal principal component analysis (PCA) to remove batch effects and were integrated using weighted-nearest neighbor (WNN) analysis, an unsupervised framework enabling integrative analysis of multiple modalities, after dimensionality reduction15. Cells were visualized using uniform manifold approximation and projection (UMAP)17. Three major populations could be readily distinguished using unsupervised clustering: pr Teff CD4+ T cells (CD4+CD154+); pr Treg CD4+ T cells (CD4+CD137+CD154−); and nr CD4+ T cells (CD4+CD154−CD137−) (Extended Data Fig. 1). Using the same approach, we subclustered pr Teff and Treg CD4+ T cell subsets into 14 subclusters based on integrated RNA and protein expression data using the WNN method and annotated based on their expression profiles (Fig. 1c,d and Supplementary Table 5): TH2a-like cells (PTGDR2high, GATA3high, IL4high, IL5high, IL13high, HPGDShigh, CD27low); conventional TH2 (TH2conv)-like cells (GATA3high, IL4high, IL5high, IL13high, IL9high); Tfh-like cells (CXCR5high); TH1 and TH17-like cells (IL17Fhigh, CCL20high, RORChigh, IFNGhigh, TBX21high); TH22 and TH17-like cells (IL22high, CCR6high); TH1 cytotoxic T lymphocyte (CTL)-like cells (THBS1high, NKG7high, GNLYhigh, GZMBhigh, GZMHhigh, PRF1high, CXCR3high); two subtypes of activated Teff (act1 and act2) cells (IFITM3high, IFITM2high, TNFSF10high, IL7Rhigh); two subtypes of pr Treg cells, that is, Treg_act1 (CCR6high, ENTPD1high) and Treg_act2 (SELPLGhigh); and four subtypes of pr naive cells (TNa_1, TNa_2, TNa_3, TNa_4).
To dissect clonotypic features among 72,491 pr CD4+ T cell phenotypes, we retrieved TCRα and TCRβ sequences from 65,032 pr CD4+ T cells with 40,897 of them having paired TCRα and TCRβ sequences. Analysis revealed distinct clonal lineages among the 14 clusters of pr CD4+ T cells, with minimal overlap between subclusters sharing similar phenotypes (Fig. 1e). Clonally expanded cells within an individual were identified based on common α-chain and β-chain complementarity-determining region 3 (CDR3) nucleotide sequences with the same nucleotide sequences of the V, DJ and C region (the most stringent way to define clonotype18), with low levels observed among pr naive cells and high levels among TH2a-like and TH1 CTL-like cells (Fig. 1f,g). Peanut stimulation upregulated CD154 on pr naive-like cells at 18 h without inducing memory transition or clonal expansion, suggesting early activation or bystander response. As these cells showed limited association with clinical outcomes (Supplementary Figs. 1 and 2), we did not focus on this population in this study.
Peanut OIT reduces TH2-related phenotype and enhance TH1 cytotoxic phenotypes
To investigate pr T cell phenotypic changes during OIT, we first checked subcluster frequencies within pr Teff memory cells and found that the frequency of TH2a-like cells among pr Teff memory cells decreased significantly (Fig. 2a; q = 0.02); the frequency of TH2conv-like, TH22 and TH17-like, and Teff-act2 cells each showed a decreasing trend (Fig. 2a; q = 0.09, q = 0.09 and q = 0.1, respectively). Notably, the frequency of TH1 CTL-like, and TH1-like and TH17-like, cells increased significantly (Fig. 2a; q = 0.02 and q = 0.02, respectively); Tfh-like cells showed a nonsignificant increase during OIT in active participants (Fig. 2a; q = 0.07). Placebo participants had no significant changes in frequency in any subcluster between BL and week 104 (Fig. 2a). Additionally, we noted that participants with PA compared to participants without PA had higher frequencies of TH2conv-like cells (Extended Data Fig. 2a; P = 0.017). The frequency of pr TH2a-like cells among pr Teff memory cells was not significantly different between groups (Extended Data Fig. 2b), probably because individuals without PA have fewer pr CD4⁺ T cells among total CD4+ T cells19 and because of the small number of participants without PA who were evaluated. In addition, TH2a-like clonotypes showed less clonal expansion in individuals without PA compared to participants with PA (Extended Data Fig. 2c).
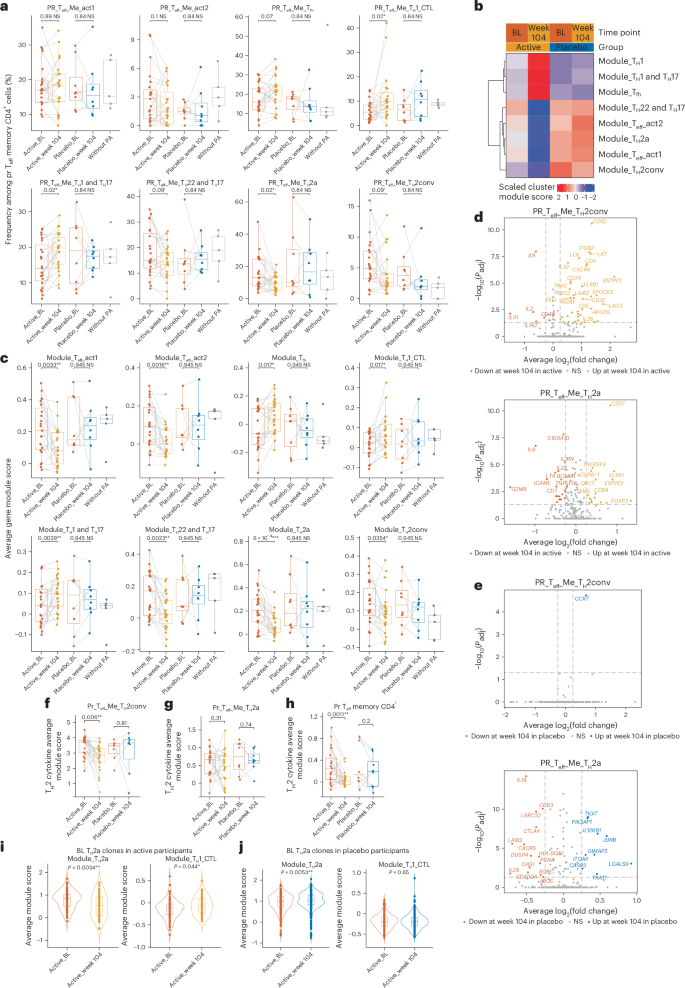
a, Box plots showing the frequency of each pr Teff memory subcluster among pr Teff memory cells in the active (n = 23) and placebo (n = 8) groups at BL and week 104, and participants without PA (n = 5). b, Heatmap showing the scaled average gene module scores of the top expressed feature genes in each subcluster in the active or placebo groups at BL and week 104. c, Box plots showing the indicated average subcluster feature gene module score of pr Teff memory cells in active participants (n = 23), placebo participants (n = 8) and participants without PA (n = 5) at BL and week 104. d, Volcano plots showing DEGs (Padj < 0.05 and average log2(fold change) > 0.25) comparing the active group at week 104 versus the BL in pr TH2conv-like (top) and pr TH2a-like (bottom) cells. e, Volcano plots showing DEGs, comparing the placebo group at week 104 with the BL in pr TH2conv-like (top) and pr TH2a-like (bottom) cells. Down, downregulated; up, upregulated. f–h, Box plots showing the average TH2 cytokine gene module scores of pr TH2conv-like (f), pr TH2a-like (g) and pr Teff memory (h) cells in the active (n = 23) and placebo (n = 8) groups at BL and week 104. i,j, Violin plots showing TH2a (left) and TH1 CTL (right) module scores of clones classified as TH2a-like cells at BL and persisting at week 104 in the active (n = 130 cells) (i) and placebo (n = 681 cells) (j) groups. In a, c and f–h, a pairwise two-sided Wilcoxon signed-rank test was used. In i and j, a two-sided Wilcoxon rank-sum test was used. In d and e, a MAST test was used. In a and c–e, multiple testing correction was performed using the Benjamini–Hochberg (false discovery rate (FDR)) method. The box plots show the median and 25th–75th percentiles; the whiskers represent 1.5 times the interquartile range (IQR); outliers are shown beyond the whiskers. All tests were two-sided. ′P < 0.1, *P < 0.05, **P < 0.01, ***P < 0.001. NS, not significant.
Source data
Peanut-specific and Ara h 2-specific IgE significantly decreased (Extended Data Fig. 3a; P = 0.00098 and P = 0.012, respectively), while the peanut-specific IgG4-to-IgE ratio and Ara h 2-specific IgG4-to-IgE ratio (Extended Data Fig. 3a; P = 2.4 × 10−7 and P = 4.8 × 10−7, respectively) each significantly increased during OIT in the active but not in the placebo group (Extended Data Fig. 3b). We examined the relationship between antibody measurements and subcluster frequencies among pr Teff memory cells at the combined BL and week 104 time points in active and placebo participants (Extended Data Fig. 3c). The frequencies of TH2conv-like and TH2a-like cells were positively correlated with peanut-specific and Ara h 2-specific IgE levels and negatively correlated with IgG4-to-IgE ratios. In contrast, TH1 CTL-like cell frequencies showed the opposite pattern, positively correlating with IgG4-to-IgE ratios and negatively correlating with IgE levels. TH1 CTL-like, and TH1-like and TH17-like, frequencies were positively correlated with each other and inversely correlated with TH2-like subsets (Extended Data Fig. 3d). The TH2conv-like, TH2a-like and TH1 CTL-like subcluster frequencies at the BL time point also showed similar patterns of correlation with each other and with the BL serological measurements (Extended Data Fig. 3e,f).
We next evaluated cluster-specific feature gene module scores in pr Teff memory CD4+ T cells to better capture both subcluster frequency and phenotype changes. Module scores reflect the expression of key subtype-defining genes (Supplementary Table 5 and Methods) and allow assessment of functional shifts even when cell frequencies remain stable. We found that during active OIT, but not placebo treatment, the module scores of TH2a-like, TH2conv-like, TH22-like and TH17-like, Teff-act1 and Teff-act2 cells significantly decreased, while TH1 CTL-like, TH1-like and TH17-like, and Tfh-like cells significantly increased in pr Teff memory cells (Fig. 2b,c). The gene module score and subcluster frequency analysis showed congruent patterns; they probably represent different but complementary views of the same underlying biological signal. Notably, although the frequency of TH2conv-like cells showed a nonsignificant downward trend, the TH2conv gene module score significantly decreased during OIT, suggesting phenotypic suppression beyond changes in frequency. Differentially expressed gene (DEG) analysis further revealed reduced expression of the type 2 cytokine interleukin-9 (IL-9) and the proliferation cytokine interleukin-2 in TH2conv-like cells during OIT in the active but not in the placebo group (Fig. 2d,e). At the individual level, TH2 cytokine module scores (IL4, IL5, IL9, IL13) in TH2conv-like cells were significantly reduced during OIT (Fig. 2f; P = 0.006) but not in the placebo group. While IL-9 expression also decreased in TH2a-like cells during OIT (Fig. 2d,e), their overall TH2 cytokine module score decrease did not reach significance (Fig. 2g). However, when considering all pr Teff memory cells, the TH2 cytokine module score significantly decreased during OIT (Fig. 2h; P = 0.0011), supporting suppression of TH2-related phenotypes beyond the decrease of cell numbers in TH2 cell subsets.
To track pr CD4+ T cell clonal frequency changes more broadly, we performed bulk TCRβ sequencing (TCRβ-seq) on PBMCs from the same participants and time points. The summed frequency of BL pr Teff memory CD4+ clones significantly decreased from BL/week 0 to week 104 in active but not placebo participants (Extended Data Fig. 4a; P = 0.011). BL TH2a and TH2conv clones also declined during active OIT, which is consistent with their role in allergy pathogenesis and treatment response (Extended Data Fig. 4b). In contrast, BL TH1 CTL-like clones did not significantly expand in the bulk data at week 104, suggesting that the increased TH1 CTL cell frequency detected in single-cell data reflects either newly generated clones during OIT or potential expansion of small BL clones below detection (Extended Data Fig. 4b).
To test another key question, whether OIT induces phenotypic shifts in T cell clones, we analyzed clones (defined by identical TRAV/TRAJ/TRBV/TRBD/TRBJ and CDR3α and CDR3β sequences) present at both BL and week 104. We detected 738 and 946 clones in active and placebo group across time points, with most clones maintaining consistent phenotypes. However, compared to the placebo group, fewer TH2a-like clones persisted as TH2a, with more shifting to TH1 CTL-like or TH1-like and TH17-like states in active participants (Extended Data Fig. 4c,d). Analysis of BL TH2a-like clones still detectable at week 104 in active participants confirmed this shift, with decreased TH2a module scores (Fig. 2i, left; P = 0.0034) and increased TH1 CTL module scores (Fig. 2i, right; P = 0.044), suggesting a transcriptional shift from a TH2a-like phenotype toward a TH1 CTL-like state induced by OIT. In contrast, BL TH2a-like clones in the placebo participants showed increased TH2a module scores (Fig. 2j, left; P = 0.0053) and no change in TH1 CTL module scores (Fig. 2j, right; P = 0.65).
Together, these data indicate that OIT attenuates pathogenic TH2a-like clonotypes and suppresses TH2-associated phenotypes in both TH2a-like and TH2conv-like cells, while promoting TH1 CTL cell frequency, TH1-associated phenotypes and also potentially inducing phenotype transitions from TH2a to TH1 cytotoxic states.
Peanut OIT leads to TH2a-like cell clonal contraction and TH1 CTL-like cell expansion
In addition to phenotype analysis, we also examined TCR clonal expansion, defining expanded clones as clonotypes presented in more than two cells in a sample (Fig. 3a). Among active participants, the proportion of expanded TH2a-like cells significantly declined during OIT, both relative to total pr Teff memory T cells (Fig. 3b; P = 0.0083) and within the TH2a-like population (Fig. 3c; P = 0.025). In contrast, clonally expanded TH1 CTL-like cells increased over time (Fig. 3b, P = 0.076; Fig. 3c, P = 0.044). These results indicate that peanut OIT drives TH2a-like clonal contraction alongside TH1 CTL-like expansion, suggesting that the relative increase in TH1 cytotoxic phenotypes is not solely the arithmetic consequence of decreases in the proportion of TH2 phenotype cells.
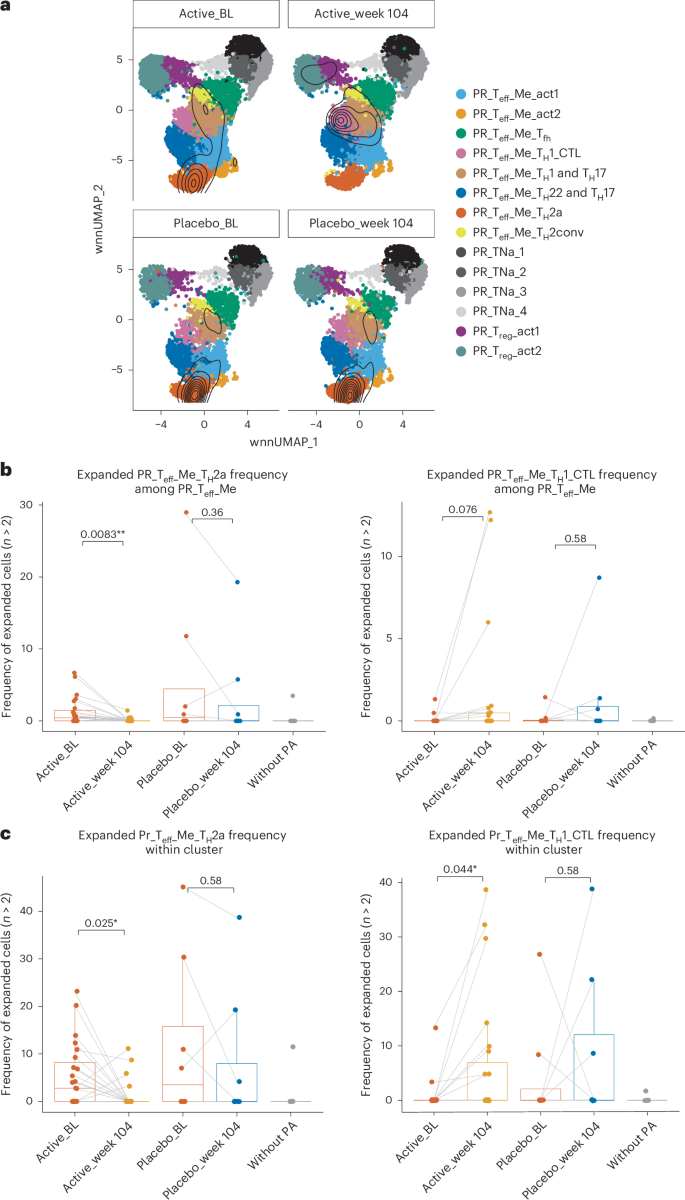
a, Overlay of the position of clonally expanded cells with a clone size larger than 2 mapped onto corresponding UMAPs for active and placebo groups at BL and week 104. b, Percentage of expanded pr TH2a-like (left) and pr TH1 CTL-like (right) cells (clonal size greater than two) among pr Teff memory cells between BL and week 104 in active (n = 23) and placebo individuals (n = 8) at BL and week 104, and participants without PA (n = 5). P values were obtained using a pairwise two-sided Wilcoxon signed-rank test. c, Percentage of expanded cells (clonal size greater than 2) within the pr TH2a-like (left) and pr TH1 CTL-like (right) clusters between BL and week 104 in active (n = 23) and placebo individuals (n = 8) at BL and week 104, and participants without PA (n = 5). In b and c, a pairwise two-sided Wilcoxon signed-rank test was used. All tests were two-sided. The box plots show the median, the 25th–75th percentiles, whiskers to 1.5 times the IQR and outliers beyond the whiskers. *P < 0.05, **P < 0.01.
Source data
Subtle phenotypic changes in pr Treg cells during OIT
Treg cells are important in immune tolerance, but we did not observe significant frequency or phenotype changes in pr Treg subclusters during OIT (Extended Data Fig. 5a–d). DEG analysis of pr Treg cells from active participants between week 104 and BL revealed only two significantly altered genes: CXCR3 (average log2(fold change) = 0.406, Padj = 0.044) was upregulated, and IER3 (average log2(fold change) = −0.468, Padj = 0.002) was downregulated (Extended Data Fig. 5e). Notably, CXCR3 expression was also higher in the active group than in the placebo group at week 104 (average log2(fold change) = −0.539, Padj = 6.04 × 10−6) (Extended Data Fig. 5f). Additionally, the frequency of BL pr Treg clones in the total PBMC repertoire remained stable across all time points (Extended Data Fig. 4a). These findings align with the hypothesis that CD137+ pr Treg cells are not primarily affected by OIT.
Distinct pr Teff memory CD4+ cells in SU and DS individuals
Next, we investigated the phenotypes associated with SU or DS outcomes in pr Teff memory CD4+ cells. While subcluster frequencies did not significantly differ between SU and DS at the individual level, we observed distinctive gene expression profiles between SU and DS participants at BL, week 104 and week 117 (Fig. 4a,b). Specifically, SU participants had lower TH2conv-like module scores at BL, lower Tfh-like module scores at week 117, higher TH22-like and TH17-like module scores at BL, and higher TH1 CTL-like module scores across all time points.
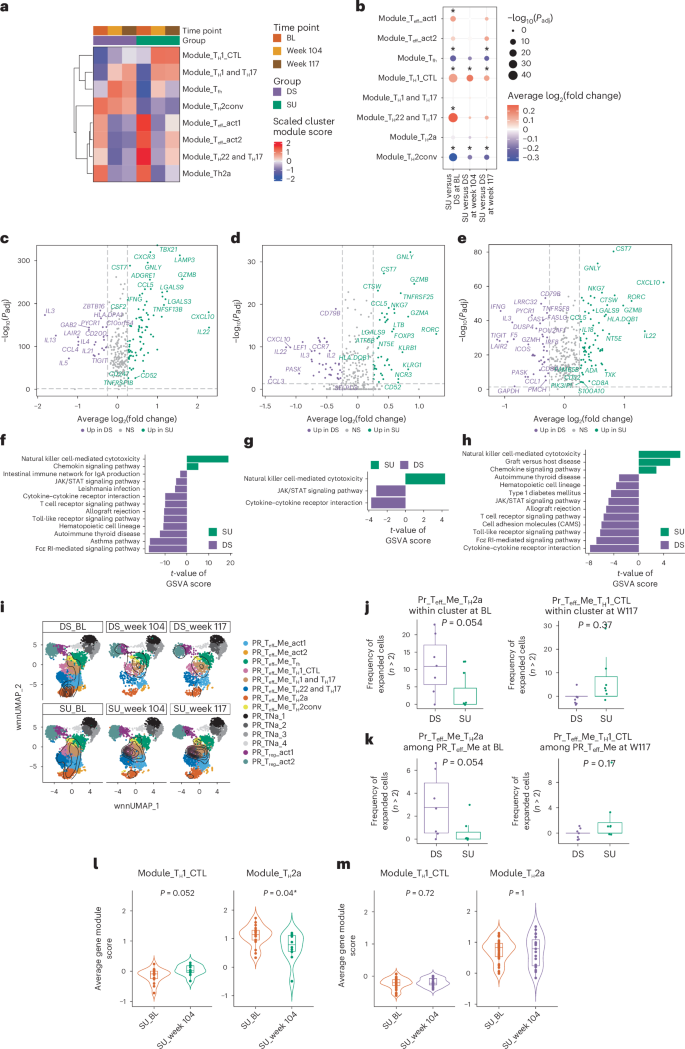
a, Heatmap showing the scaled average gene module scores of the top expressed feature genes in each cell subcluster in the SU or DS group at BL, week 104 and week 117. b, Comparison of the average gene module scores of the top expressed feature genes in each cell subcluster between the SU and DS groups at BL, week 104 and week 117 using the MAST test. P values were Benjamini–Hochberg (FDR) corrected. Dot size represents −log10(Padj); color represents the average log2(fold change). c–e, Volcano plots showing DEGs comparing pr Teff memory CD4+ T cells in the SU and DS groups at BL (c), week 104 (d) and week 117 (e), obtained using the MAST test. P values were FDR adjusted. f–h, Differences in pathway activities scored using GSVA in SU versus DS at BL (f), week 104 (g) and week 117 (h). The pathways shown have top-ranked absolute t-values obtained using a linear model with an FDR-adjusted P < 0.01. i, Overlay of the position of clonally expanded cells with a clone size larger than 2 mapped onto corresponding UMAPs for the SU and DS groups at BL, week 104 and week 117. j, Comparison of the percentage of expanded cells (clonal size greater than 2) within the pr TH2a-like cluster at BL (left) and pr TH1 CTL-like cells at week 117 (right) between SU (n = 7) and DS (n = 7) participants. P values were obtained using a two-sided Wilcoxon rank-sum test. k, Comparison of the percentage of the expanded pr TH2a-like cluster at BL (left) and pr TH1 CTL-like cells at week 117 (right) (clonal size greater than 2) among total pr Teff memory cells between SU (n = 7) and DS (n = 7) participants. l,m, Violin plots showing the TH1 CTL (left) and TH2a (right) module scores of clonotypes classified as TH2a-like cells at BL and still present at week 104 in the SU (n = 27 cells) (l) and DS (n = 53 cells) (m) groups. In j–m, a two-sided Wilcoxon rank-sum test was used. In c–e, a MAST test was used. In b–h, multiple testing correction was performed using the Benjamini–Hochberg (FDR) method. All tests were two-sided. *P < 0.05.
Source data
DEG analysis on pr Teff memory CD4+ cells between SU and DS participants at BL, week 104 and week 117 (Fig. 4c–e) further revealed that the key feature genes related to cytotoxicity phenotype (for example, PRF1, GNLY, NKG7, GZMB, CST7) were higher in the SU than in the DS group at BL and week 117; TH2-related genes (for example, IL13, IL4, IL5) were higher in the SU than in the DS group at BL. Gene set variation analysis (GSVA)20 confirmed upregulation of pathways related to cytotoxicity signaling in pr Teff memory CD4+ cells in SU participants at all three time points, while pathways related to asthma were lower in SU participants at BL compared to DS participants (Fig. 4f–h). To evaluate whether differences in gene expression between SU and DS participants differed between OIT time points, we carried out correlation analysis of gene expression effect sizes (average log2(fold change)) between SU and DS participants for each pair of time points. This analysis revealed that SU versus DS differences in gene expression were more highly correlated between BL and week 117 (R = 0.62) than between BL and week 104 (R = 0.46) or between week 104 and week 117 (R = 0.50) (Extended Data Fig. 6), suggesting that gene expression differences at week 104 after continuous allergen exposure are of limited duration.
TCR clonal analysis revealed higher clonal expansion in the TH1 CTL-like subcluster in pooled SU participant data compared to pooled DS participant data at week 104 and week 117, and in the TH2a subcluster in the DS group than in the SU group at BL (Fig. 4i). In the individual participant-level analysis, the frequency of expanded pr TH2a-like cells within the TH2a cluster or among pr Teff memory cells was lower in SU compared to DS participants at BL (Fig. 4j, P = 0.054; Fig. 4k, P = 0.054), while the frequency of expanded pr TH1 CTL-like cells within the pr TH1 CTL cluster or among pr Teff memory cells showed a nonsignificant trend toward higher values in SU compared to DS participants at week 117 (Fig. 4j, P = 0.37; Fig. 4k, P = 0.17).
In SU participants, TH2a-like clonotypes persisting from BL to week 104 showed a significant decrease in TH2a module score (Fig. 4l; P = 0.04) and a trend toward increased TH1 CTL module score (Fig. 4l; P = 0.052), with no changes in DS participants (Fig. 4m). DEG analysis of TH2a-like cells between SU and DS participants revealed no changes at week 104, but at BL and week 117. SU cells upregulated cytotoxic/TH1 CTL-related genes (for example, NKG7, GNLY, PRF1, TBX21), while DS cells expressed higher TH2 cytokines (for example, IL4, IL5, IL13) (Extended Data Fig. 7a,b). DEG patterns between SU and DS mirrored differences between the TH1 CTL-like and TH2a-like subsets (Extended Data Fig. 7c,d), indicating a phenotypic shift rather than a difference in subset abundance. Supporting this, we found that the average TH1 CTL module score in TH2a-like cells was consistently higher in SU participants, reaching statistical significance at week 117 at the individual level (Extended Data Fig. 7e; P = 0.022) and trending upward over time in SU but not in DS participants. These results suggest that a greater potential for TH2a-like cells to adopt a TH1 CTL-like phenotype may contribute to the achievement of SU.
Lower BL TH2 gene signatures are associated with higher likelihood of SU outcome
Given the higher prevalence of the TH2-related phenotype in DS compared to SU participants, we calculated the average module scores of TH2-related cytokine gene sets and Kyoto Encyclopedia of Genes and Genomes asthma pathway gene sets (CD40, CD40LG, FCER1G, IL13, IL3, IL4, IL5, TNF, shown in Fig. 5a) in pr Teff cells for each participant at BL, week 104 and week 117. Both the TH2 cytokine and asthma pathway gene module were relatively elevated in DS participants at BL (Fig. 5b; P = 0.073 and P = 0.053, respectively). These features declined more noticeably in DS participants over time, potentially because of their initially higher levels, and was more comparable between SU and DS participants by week 104, with a modest trend toward remaining slightly higher in the DS group at week 117. Receiver operating characteristic (ROC) analysis showed that the TH2 cytokine module score at BL had the highest potential for distinguishing SU from DS (Fig. 5c; area under the curve (AUC) = 0.7959, P = 0.0639) and was significantly correlated with the final clinical outcome (peanut cumulative tolerated dose (CTD) during food challenge at week 117) (Fig. 5d; R = 0.57, P = 0.033). Similarly, the asthma pathway module score at BL had the highest potential for distinguishing SU from DS after OIT (Fig. 5e; AUC = 0.8163, P = 0.0476) and was significantly correlated with the final clinical outcome (Fig. 5f; R = 0.68, P = 0.007). Together, these results highlight that BL TH2-related activity is associated with SU outcomes.
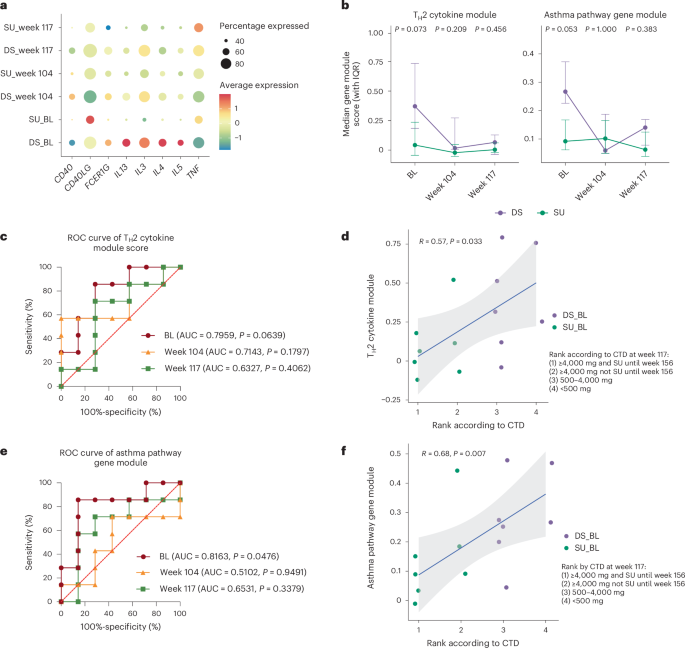
a, Dot plot displaying the average expression and frequency of gene-expressing cells for each TH2-related gene among pr Teff memory cells in the SU and DS groups at each time point. b, Line plot showing the longitudinal changes in the indicated gene module scores in pr Teff memory cells from SU (n = 7) and DS (n = 7) participants at BL, week 104 and week 117. The dots represent the medians of each individual in each group; the error bars indicate the IQR. P values were calculated using a two-sided Wilcoxon rank-sum test to compare SU (n = 7) and DS (n = 7) individuals at each time point. c, ROC analysis for the prediction of clinical outcome after OIT (SU or DS), based on the module scores of TH2 cytokines at BL, week 104 and week 117. P values were obtained from the ROC analysis without adjustment. d, Scatter plots showing the Spearman’s rank correlation between the module scores of TH2 cytokines at BL and the rank of clinical outcome according to CTD at week 117 (CTD ranking: 1. ≥4,000 mg and remained SU until week 156; 2. ≥4,000 mg but did not remain SU until week 156; 3. 500–4,000 mg; 4. <500 mg). The trend line from the linear regression is shown, with the shaded band showing the 95% confidence interval (CI) based on the standard error. e, ROC analysis for the prediction of clinical outcome after OIT (SU or DS) based on the asthma pathway gene module scores at BL, week 104 and week 117. P values were obtained from the ROC analysis without adjustment. f, Scatter plot showing the Spearman’s rank correlation between the asthma pathway gene module scores at BL and the rank of clinical outcome according to CTD at week 117 (rank as defined above). The trend line from the linear regression is shown, with the shaded band showing the 95% CI based on the standard error. All tests were two-sided.
Source data
Higher cytotoxicity gene signatures after OIT are linked to higher likelihood of SU outcome
In addition to the TH2-related phenotype, we also found that pr TH1 cytotoxicity-related features that changed during OIT could also contribute to clinical outcome. We calculated the average module scores of CD4 cytotoxicity-related genes (GZMB, PRF1, GNLY, KLRG1, NKG7, CST7, GZMH, CX3CR1; shown in Fig. 6a)21 and pr TH1 CTL feature genes in pr Teff memory cells for each participant at BL, week 104 and week 117, and found that the CD4 cytotoxicity module showed a clearer upward trend over time in SU participants, ultimately reaching significance at week 117 (Fig. 6b, left; P = 0.017). The pr TH1 CTL feature gene module score showed a consistent trend of higher values in SU compared to DS participants across all time points; it most closely approached significance at week 117 (Fig. 5b, right; P = 0.053). These findings suggest that immune dynamics over time are relevant to the outcome. In addition, the ROC analyses showed the CD4 cytotoxicity module score at week 117 as having the highest potential for distinguishing SU from DS (Fig. 6c; AUC = 0.8776, P = 0.0181) and being significantly correlated with clinical outcome (Fig. 6d; R = −0.59, P = 0.025). Similarly, the TH1 CTL feature gene module score at week 117 had the highest potential for distinguishing SU from DS after OIT (Fig. 6e; AUC = 0.8163, P = 0.0476) and was significantly correlated with the final clinical outcome (Fig. 6f; R = −0.57, P = 0.033). Together, these results highlight that the cytotoxicity-related phenotype after OIT is associated with SU outcomes.
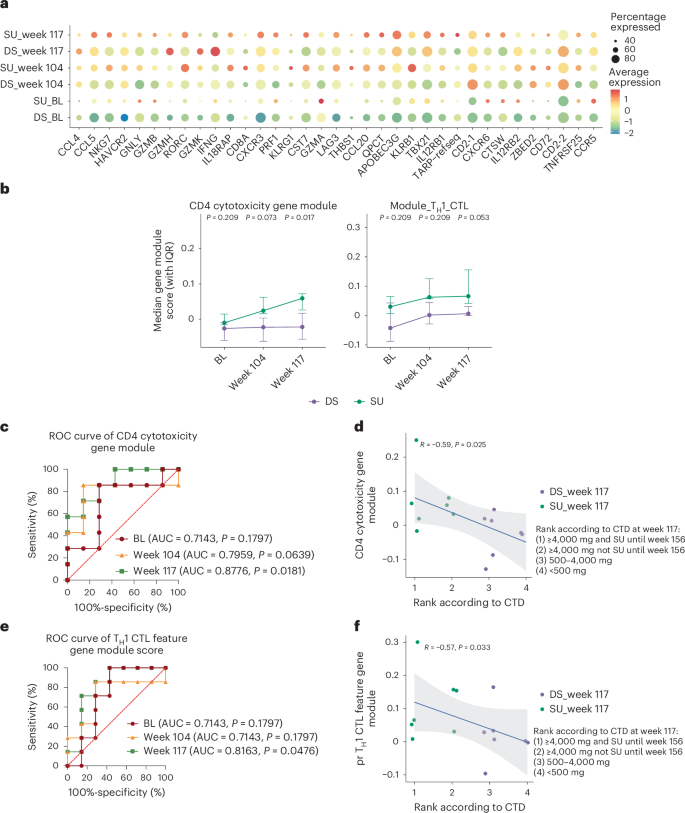
a, Dot plot displaying the average expression and frequency of gene-expressing cells for each CD4 cytotoxicity-related gene among pr Teff memory cells in the SU and DS groups at each time point. b, Line plot showing longitudinal changes in the indicated gene module scores in pr Teff memory cells from SU (n = 7) and DS (n = 7) participants at BL, week 104 and week 117. The dots represent the medians of each individual in each group; the error bars indicate the IQR. P values were calculated using a two-sided Wilcoxon rank-sum test to compare SU (n = 7) and DS (n = 7) individuals at each time point. c, ROC analysis for the prediction of clinical outcome after OIT (SU or DS) based on the CD4 cytotoxicity-related gene module scores at BL, week 104 and week 117. P values were obtained from the ROC analysis without adjustment. d, Scatter plots showing the Spearman’s rank correlation between the module scores of CD4 cytotoxicity-related genes at week 117 and the ranking of clinical outcome according to CTD at week 117. The trend line from the linear regression is shown, with the shaded band showing the 95% CI based on the standard error. e, ROC analysis for the prediction of clinical outcome after OIT (SU or DS) based on the module scores of TH1 CTL-like cluster feature genes at BL, week 104 and week 117. P values were obtained from the ROC analysis without adjustment. f, Scatter plots showing the Spearman’s rank correlation between the module scores of TH1 CTL-like cluster feature genes at week 117 and the ranking of clinical outcome according to CTD at week 117. The trend line from the linear regression is shown, with the shaded band showing the 95% CI based on the standard error. All tests were two-sided.
Source data
Higher CD39 expression on pr Treg cells is associated with SU
Although the subcluster frequency and cluster feature gene module score of pr Treg cells were not significantly different between SU and DS (Extended Data Fig. 8a,b), there were immune-related DEGs in pr Treg cells between SU and DS at week 117, week 104 and BL (Fig. 7a). ENTPD1 (encoding CD39), which has been primarily described as a Treg marker22,23 and is crucial in terms of immunosuppressive functions22,23, was highly enriched in pr Treg than in pr Teff (Fig. 7b) cells. ENTPD1 (CD39) expression was significantly higher in pr Treg cells in SU participants than in DS participants at BL and week 117 (Fig. 7c; average log2(fold change) = 1.005, Padj = 5.75 × 10−12 and average log2(fold change) = 1.84, Padj = 2.38 × 10−22, respectively). Both the RNA and protein levels of ENTPD1 (CD39) in pr Treg cells at the individual level at week 117 of the OIT could distinguish SU and DS participants (Fig. 7d; AUC = 0.9388 and AUC = 0.9592). The frequency of ENTPD1-expressing cells and CD39+ cells (Supplementary Fig. 3) among pr Treg cells was significantly higher in SU than DS participants at week 117 (Fig. 7e,f; P = 0.011 and P = 0.0072). In addition, both the RNA and protein levels of ENTPD1 (CD39) at week 117 in pr Treg cells was significantly correlated with the outcome of peanut CTD during the week 117 food challenge (Fig. 7g,h; R = −0.63, P = 0.015; R = −0.76, P = 0.0015). To explore potential suppressive mechanisms of CD39⁺ pr Treg cells, we compared their gene expression with CD39− pr Treg cells (Fig. 7i). CD39⁺ pr Treg cells exhibited elevated expression of LGALS1 (galectin-1), IL3RA (interleukin-3Rα) and CD27, supporting Treg cell stability, survival and suppressive function24,25,26,27,28,29, while LIF and LAG3, which are involved in interleukin-10-mediated and STAT3/RORγt-mediated suppression30,31, were lower, suggesting it may rely on alternative suppressive pathways.
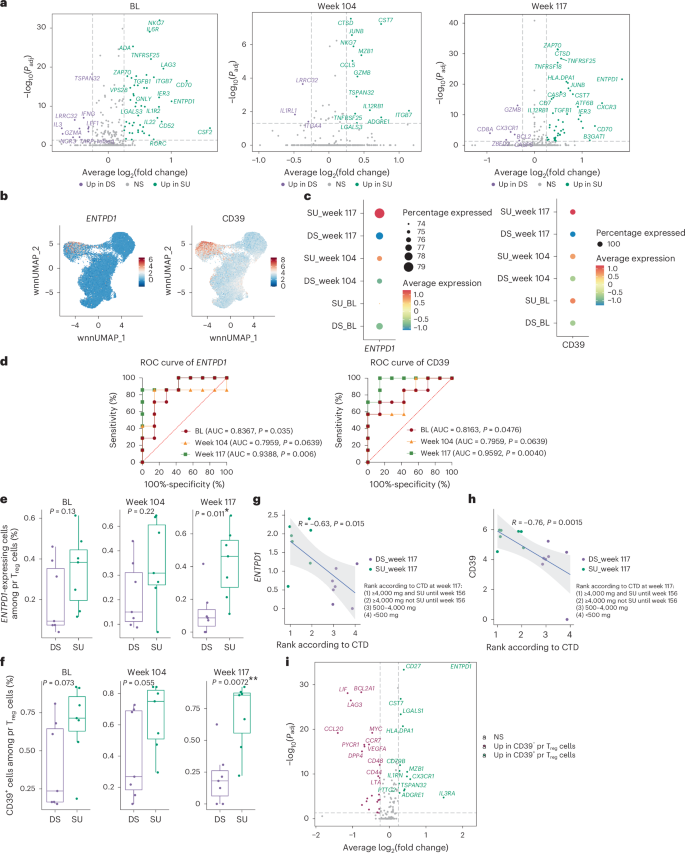
a, Volcano plots showing DEGs, comparing pr Treg cells in the SU and DS groups at BL (left), week 104 (middle) and week 117 (right), obtained using the MAST test. P values were FDR adjusted. b, UMAPs showing the expression level of ENTPD1 (encodes CD39) and CD39 (based on scAb-seq data) in pr CD4+ T cells. c, Dot plot displaying the average expression and frequency of gene-expressing cells for ENTPD1 and CD39 expression among pr Treg cells for each group and time point. d, ROC analysis for the prediction of clinical outcome after OIT (SU or DS), based on the expression level of ENTPD1 (left) and CD39 (right) at BL, week 104 and week 117. P values were obtained from the ROC analysis without adjustment. e,f, Box plots showing the difference of the percentage of ENTPD1-expressing cells (cells with an ENTPD1 raw count greater than 0) (e) and CD39+ cells (cells with an anti-CD39 antibody log-normalized and scaled expression greater than 5; Supplementary Fig. 3) (f) among pr Treg cells between SU and DS participants at BL, week 104 and week 117 (SU: n = 7; DS: n = 7). P values were obtained using a two-sided Wilcoxon rank-sum test. The box plots show the median, the 25th–75th percentiles, the whiskers to 1.5 times the IQR and outliers beyond the whiskers. g,h, Scatter plots showing the Spearman’s rank correlation between the expression level of ENTPD1 (g) and CD39 (h) at week 117, and the ranking of clinical outcome according to CTD at week 117. The trend line from the linear regression is shown, with the shaded band showing the 95% CI based on the standard error. i, Volcano plot showing DEGs, comparing CD39+ and CD39− pr Treg cells, obtained using the MAST test. P values were FDR adjusted. *P < 0.05, **P < 0.01. All tests were two-sided.
Source data
Pr clones were detected in the TCRβ repertoires from the gastrointestinal tract
To assess mucosal T cell changes, we performed bulk TCRβ sequencing on gastrointestinal (GI) biopsies from POISED participants at week 0 (n = 7) and week 104 (n = 4) (Supplementary Table 2) and identified 766 unique clones (2,471 instances) matching pr T cells from the single-cell data (Methods), broadly distributed across the upper GI tract (Supplementary Fig. 4a–d). Overall pr clone frequencies were stable, but pr Treg cell clones trended upward in the proximal esophagus, stomach and duodenum, and TH1 CTL-like clones in the duodenum (Supplementary Fig. 4e,f). Clones with a TH2a-like phenotype trended upward in the stomach, while TH2conv-like clones trended downward (Supplementary Fig. 4f), suggesting distinct dynamics in GI tissue compared to blood, or different phenotypes for the clones in the GI tract.
Taken together, these findings highlight coordinated alterations in pr CD4+ T cell populations during OIT. A summary of the key immune shifts associated with SU is shown in Fig. 8.
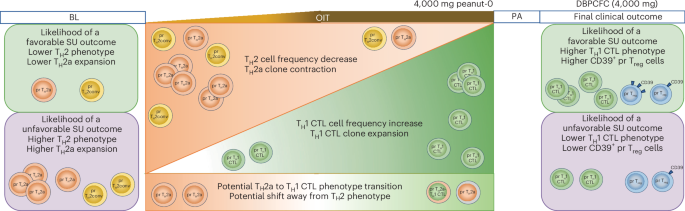
During OIT, TH2 cell frequency and TH2-associated phenotypes decrease with contraction of TH2a clones, whereas TH1 CTL cell frequency and clonal expansion increase. Higher baseline TH2 phenotype and expansion of TH2a clones are associated with a lower likelihood of achieving SU. In contrast, elevated TH1 CTL-related phenotypes and higher CD39 expression in pr Treg cells following OIT are associated with a favorable SU outcome.
link